Importance
- The curse of the dimensionality
- When Machine Learning problems involve thousands or even millions of features for each training instance
- all these features make training extremely slow
- also make it much harder to find a good solution
- high-dimensional datasets are at risk of being very sparse
- most training instances are likely to be far away from each other
- a new instance will likely be far away from any training instance
- making predictions much less reliable than in lower dimensions, since they will be based on much larger extrapolations
- Benefits of dimensionality reduction
- speed up training
- useful for visualization
Main Approaches
- Projection
- the main idea is to project all training instances to a much lower-dimensional subspace of the high-dimensional space
- Drawbacks
- Can’t use when the subspace twist and turn
- e.g. Swiss roll toy dataset
- Manifold learning
- Manifold: a d-dimensional manifold is a part of an n-dimensional space (where d < n) that locally resembles a d-dimensional hyperplane.
- Many dimensionality reduction algorithms work by modeling the manifold on which the training instances lie; this is called Manifold Learning.
- Manifold Assumption
- most real-world high-dimensional datasets lie close to a much lower-dimensional manifold
- Accompanied implicit assumption
- the task at hand will be simpler if expressed in the lower-dimensional space of the manifold.
- This one does not always hold
PCA
PCA Idea
- Main idea
- identifies the hyperplane that lies closest to the data, and then it projects the data onto it
- rotate the axis system in such a way that each axis represents a PC
- select the axis that preserves the maximum amount of variance, as it will most likely lose less information than the other projections
- Notes:
- the direction of the unit vectors returned by PCA is not stable
- if you perturb the training set slightly and run PCA again, the unit vectors may point in the opposite direction as the original vectors
- How to find PCs
- SVD
- $X=U\Sigma V^T$
- $\Sigma$ contains explained variance for each PC
- $V$ contains the unit vectors (PCs)
- Need to have centered data
- Scikit-Learn’s PCA classes take care of centering the data for you
- SVD
- How to choose the right number of dimensions
- plot the explained variance as a function of the number of dimensions.
- There will usually be an elbow in the curve, where the explained variance stops growing fast.
-
Implementation
1 2 3 4 5 6 7
# pca pca = PCA(n_components=0.95) X_reduced = pca.fit_transform(X_train) # reconstruction after compression pca = PCA(n_components = 154) X_reduced = pca.fit_transform(X_train) X_recovered = pca.inverse_transform(X_reduced)
Randomized PCA
-
Difference
- Use a stochastic algorithm called Randomized PCA that quickly finds an approximation of the first d principal components
- Its computational complexity is $O(m × d^2) +O(d^3)$, instead of $O(m × n^2) +O(n^3)$ for the full SVD approach,
- dramatically faster than full SVD when d is much smaller than n
-
Implementation
rnd_pca = PCA(n_components=154, svd_solver="randomized")- By default,
svd_solveris actually set to “auto”- Scikit-Learn automatically uses the randomized PCA algorithm if m or n is greater than 500 and d is less than 80% of m or n, or else it uses the full SVD approach.
- By default,
Incremental PCA
-
problem
- PCA and randomized PCA require the whole training set to fit in memory in order for the algorithm to run.
-
Idea of IPCA
- split the training set into mini-batches
- feed an IPCA algorithm one mini-batch at a time
-
Useful for large training sets and for applying PCA online
-
Implementation :star:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23
# method 1 from sklearn.decomposition import IncrementalPCA n_batches = 100 inc_pca = IncrementalPCA(n_components=154) for X_batch in np.array_split(X_train,n_batches): inc_pca.partial_fit(X_batch) X_reduced = inc_pca.transform(X_train) # method 2: use memmap class ## write data into memmap structure filename = "my_mnist.data" m, n = X_train.shape X_mm = np.memmap(filename, dtype='float32', mode='write', shape=(m, n)) X_mm[:] = X_train ## read X_mm = np.memmap(filename, dtype="float32", mode="readonly", shape=(m, n)) batch_size = m // n_batches inc_pca = IncrementalPCA(n_components=154, batch_size=batch_size) inc_pca.fit(X_mm)
Kernel PCA
-
Kernel trick
- a mathematical technique that implicitly maps instances into a very high-dimensional space (called the feature space)
- a linear decision boundary in the high-dimensional feature space corresponds to a complex nonlinear decision boundary in the original space.
-
Benefits of kPCA
- It is often good at preserving clusters of instances after projection
- sometimes even unrolling datasets that lie close to a twisted manifold
-
Implementation
1 2 3
from sklearn.decomposition import KernelPCA rbf_pca = KernelPCA(n_components = 2, kernel="rbf", gamma=0.04) X_reduced = rbf_pca.fit_transform(X)
-
example of different kernels
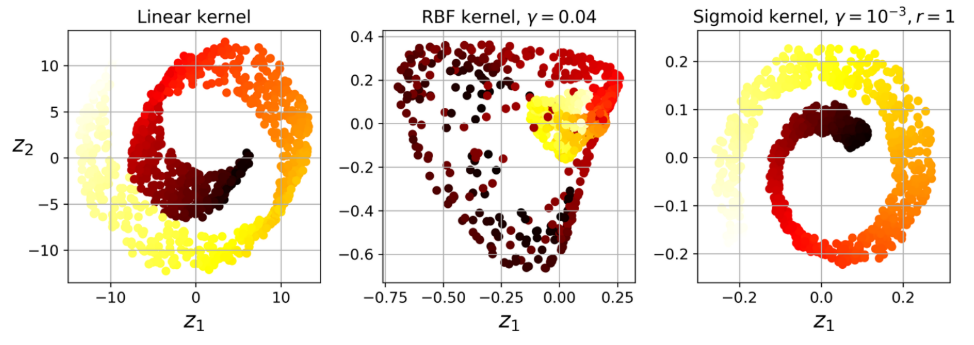
Other techniques
Locally Linear Embedding (LLE)
- nonlinear dimensionality reduction (NLDR) technique
- a Manifold Learning technique that does not rely on projections
Main idea
- first measuring how each training instance linearly relates to its closest neighbors
- then looking for a low-dimensional representation of the training set where these local relationships are best preserved.
Pros and Cons
- Pro
- good at unrolling twisted manifolds, especially when there is not too much noise.
- Con
- scale poorly to very large datasets
Implementation
1
2
3
from sklearn.manifold import LocallyLinearEmbedding
lle = LocallyLinearEmbedding(n_components=2, n_neighbors=10)
X_reduced = lle.fit_transform(X)
Other techniques
- Random Projections
- Multidimensional Scaling (MDS)
- Isomap
- t-Distributed Stochastic Neighbor Embedding (t-SNE)
- Linear Discriminant Analysis (LDA)
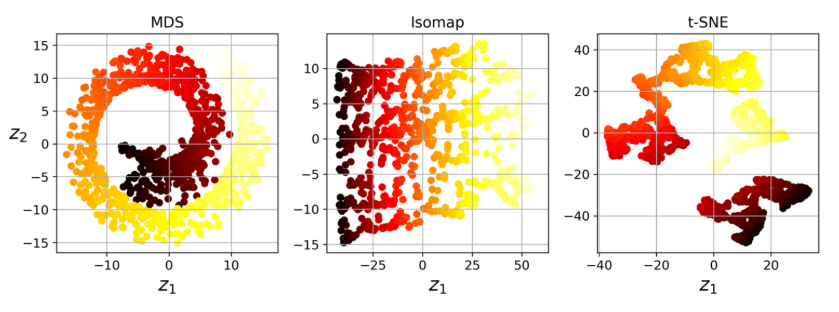
-
Previous
Machine Learning: Ensemble Learning and Random Forest -
Next
Machine Learning: Unsupervised Learning